|
This is a copy of the entrant's original web pages. The original web site may be found at
http://wwwcg.in.tum.de/vis-contest.

tum.3D wins the
IEEE Visualization Contest 2005!
Our entry to the IEEE 2005 Visualization Contest, held at the worlds largest and most renowned
scientific conference on visualization, has been ranked 1st place by an expert jury.
"All you need is ... - Particles!"
Jens Schneider,
Polina Kondratieva,
Jens Krüger,
Rüdiger Westermann
Contents
PDF: low quality [1.3MB], high quality [6.2MB]
Video: wmv, low quality [23MB], wmv, high quality [50MB]
Images/Storyboard:
New Orleans #1,
New Orleans #2,
DT-MRI Brain #1,
DT-MRI Brain #2
Downloads
Particle Engine and datasets
Contest Entry Visualization System
We visualize the given data by means of our GPU-based particle engine.
Designed for interactive visualization of steady 3D flow fields, it achieves its high frame rates by
exploiting features of recent graphics processing units (GPU) for particle advection.
Particle attributes and the velocity field are kept in graphics memory, avoiding any transfer
across the bus for rendering. This allows for rendering of millions of particles at interactive rates.
Recently, the particle engine was extended by a DT-MRI visualization module.
Entries of a symmetric, real-valued diffusion tensor are stored in 3D float-textures. For each particle,
the tensor is tri-linearly interpolated from neighbouring entries. The result is then used to either advect
particles along the first eigenvector, or to deflect them.
The user can choose from a multitude of visualization methods, such as plain particles, oriented texture
splats, shaded lines, stream balls, or stream ribbons. More context is provided by further augmenting the
result with volume rendering or triangle meshes. Consequently, rendering methods can be mapped to different
modalities without need to resort to other tools.
The entire application is based on DirectX Techniques and can be extended very easily, making it a flexible
and powerful visualization tool.
This entry is part of the
Research Project "Virtual Windtunnel"
and of the DFG SPP 1147.
It is partially funded by the Deutsche Forschungsgemeinschaft.
J. Krüger, P. Kipfer, P. Kondratieva, R. Westermann
"A Particle System for Interactive Visualization of 3D Flows"
IEEE Transactions on Visualization and Computer Graphics Vol. 11, No. 5
P. Kondratieva, J. Krüger, R. Westermann,
"The Application of GPU Particle Tracing to Diffusion Tensor Field Visualization",
IEEE Visualization 2005
Task 1: Interactive Exploration
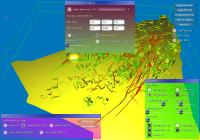
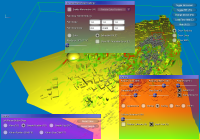
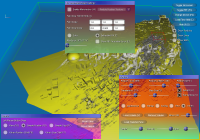
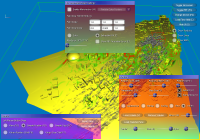
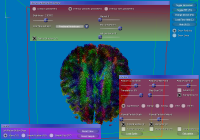
From left to right:
a) GUI for line setup, showing stream ribbons and a Lambda2 volume of the data,
b) particle setup, showing the combination of volume rendering and depth-sorted particles,
c) volume rendering setup, modifying the transfer function of the previous image,
d) more particle settings,
e) DTI parameters
Performance:
All measurements where done on a Pentium4 2.4GHz, 1 GB RAM, nVidia GeForce 6800 Ultra.
Viewport size was 1280x1024. The New Orleans data set was cropped and resampled to 512x512x64xfloat4
as discussed in the pdf. All timings use either particles of 7x7 pixels or lines/ribbons of 300
integration steps length (few, long lines are more expensive than many, short ones). In both cases
RK32 integration was used. The DTI data set has full 256x256x24 resolution as described in the pdf.
Deflection was used for particle propagation.
64K (256K) plain particles 171fps (70fps)
64K (256K) oriented texture splats 55fps (15fps)
64K (256K) depth-sorted particles 25fps ( 7fps)
128 (256) shaded lines 27fps (25fps)
128 (256) stream ribbons 17fps (16fps)
300 (500) slices volume 71fps (45fps)
64 (128) DTI stream tubes 13fps (11fps)
64K (256K) DTI oriented splats 30fps ( 8fps)
Navigation:
The user navigates either by mouse and keyboard, or, more intuitively, using a spacemouse or joystick.
A rectangular, particle-emitting �probe� of arbitrary size and position can be controlled, which, combined
with interactive frame rates, makes data exploration a highly intuitive process. The engine also includes
a GPU-based volume renderer with all standard features such as clip-planes and transfer-functions. It is
worth noting that multiple transparencies can be rendered correctly by performing a depth sort directly on
the GPU. This enables the creation of particle transfer-functions that map particle age to rgba values.
Parameters:
User-modifiable parameters fall into three classes: Primitives, Colors, Appearance. The user may choose
among primitives such as particles, lines or volumes. Particles are available in numerous appearances
(they can have the shape of any mesh, they are actually impostors), and can be oriented to better reflect
the direction of the flow. Line appearances include shaded lines, stream balls, and stream ribbons with
optional curl enhancement.
Task 2: Static Presentation - New Orleans Wind Field
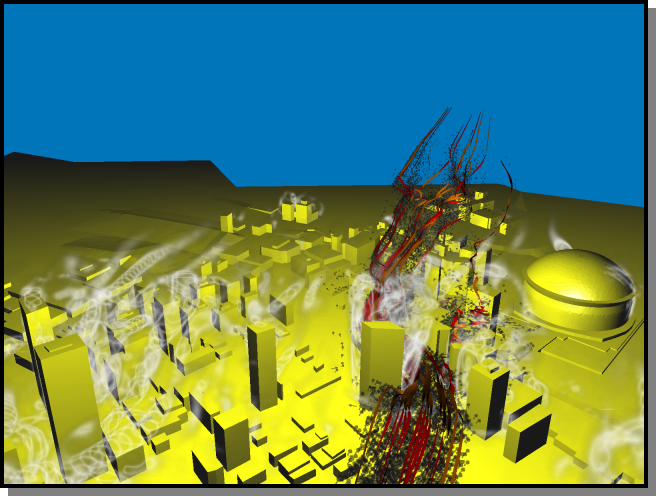 This region around the Louisiana Superdome was visualized using stream-ribbons and oriented particles. The cloud-like volumes
behind the buildings enclose vortex cores that were segmented using the lambda2 criterion (see pdf).
This region around the Louisiana Superdome was visualized using stream-ribbons and oriented particles. The cloud-like volumes
behind the buildings enclose vortex cores that were segmented using the lambda2 criterion (see pdf).
Left of the particle-stream, this volume greatly assists the user in identifying turbulent regions and probing the field accordingly.
Moreover it carries information by itself. As can be seen between the trapezoid building and the stream-ribbons in the foreground,
cores are similar to an eye of a storm. The particles whirling around them only enter them rarely. In the presence of an airborne
contaminant, a good chance for a safe place would be where vortex core and building meet.
Stream-ribbons help to identify the global appearance of the field, including its curl. Behind the first row of buildings to the
right is a highly turbulent vortical structure, which cannot be resolved sufficiently by the volume or the particles: The volume
needs to be transparent enough to allow for an unobstructed view on the structures in the back, while particles in a static image
are pointing in seemingly arbitrary directions. However, important local information about the orientation of the field is conveyed
by particles.
The triangle mesh provides the necessary context to fully understand the field.
Visualizations that do not mix rendering primitives will most likely fail to fully communicate the field in this region.
This visualization renders at ca. 22fps on our system described above.
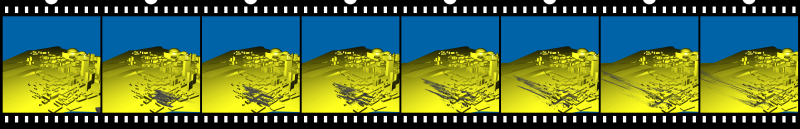

In this sequence, the particle source is located at Poydras Street Wharf, Riverwalk Marketplace, and has the approximate size of the freighter Bright Field (735x125x50ft).
Particles are emitted in a single burst to mimick the impact, and are then advected with the windfield. Note that while some particles are transported away quickly,
others are captured in vortices and stay for a long time in the same region.
This visualization renders at ca. 165fps on our system as described above.
[TOP]
Task 2: Static Presentation - DT-MRI Brain
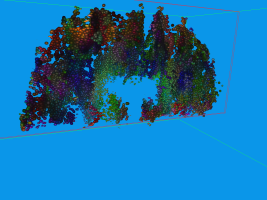
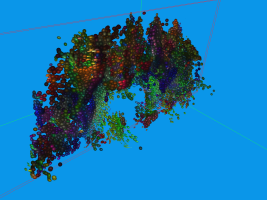
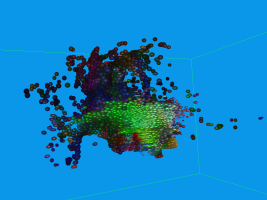
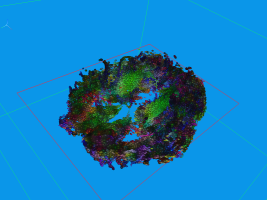




Usually DTI brain data set are visualized by glyphs or stream-tubes. However, both methods do not
communicate the fact that diffusion is a temporal process very well. In contrast, we can advect
aligned ellipsoids, conveying the movement of water in the brain a lot better. This, of course is
hard to show in static images, and consequently the images may look similar to those generated with
standard approaches.
In contrast to other approaches our ellipsoids are still advected, making it, due to varying particle
density, possible to convey both global and local structures of the underlying tensor field. This is
not possible if they are equidistantly spaced across the domain. Advection proceeds either along the
largest eigenvector of the tensor or along the most likely diffusion direction. Removing particles at
positions that do not exhibit anisotropy, the connectivity between the different regions of the brain
can be explored and distinguished. The user can choose the color scheme from standard models such as
Cl,Cs,Cp or FA.
Stream-tubes are usually better associated with the fibrous structures present in the brain tissue.
The color mapping schemes are the very same as for ellipsoids. The user explores the data by
adjusting size and position of a seeding region. This intuitive method allows for quick navigation
and efficient recognition of brain structures within the given volume.
The above sequence starts by seeding particles in a single coronal slice. After some advection steps,
the next picture was taken, where the superior longitudinal fasciculus (green) can be easily made out
along the Y-axis. Other structuees such as the corona radiata and U-shaped fibers can also be easily
found in this image, though the data set contains a considerable amount of noise (see pdf). Finally
the fourth image depicts an axial slice.
This visualization renders at ca. 15fps on our system as described above.
Another visualization using stream tubes can be found below.
[TOP]
Task 3: Data Specific Tasks
Most of the data specific tasks for the New Orleans data set were already covered in the previous section, namely the identification of vortices and a
sequence showing the time evolution of an airborne pollutant. Over that, various other visualizations were performed, some of which we want to present here.
Visualization Performed 1
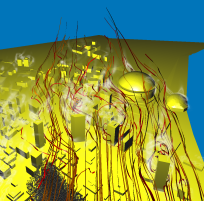
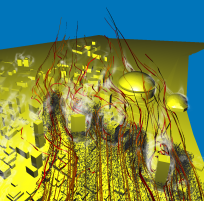
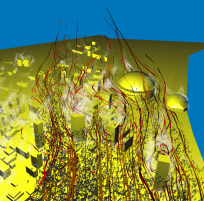
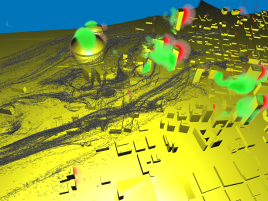



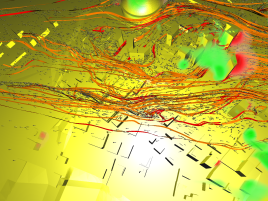

After having already performed the visualization for task 2, we were interested in getting a broader view of the flow close to the Superdome. Again, we proceeded
very similar as before, but this time we took a very large particle source and simulated a sudden burst dispersing material into the air. Stream-ribbons where used
to provide the viewer with the necessary context. It turns out that the front of particles is advancing (first image), quickly hits the building (second image) and splits
into several streams (third image). In the fourth image, we displayed the energy volume built from the energy values present in the original data. It turns out that the
local pressure maximum that one would expect in front of buildings can be highlighted using a simple red-to-green transfer function. We think that the signed energy values
hold the difference between dynamic (i.e. kinetic) and static (i.e. pressure) energy. Negative signs correspond to higher static energy, while positive values corespond to
high velocity. Clearly atop of the roof of the Superdome there is a high velocity, as the roof is curved. Behind buildings areas of high velocities correspond to vortex
cores and are a first guess for safe places, as can be seen behind the trapezoid shaped building. In the last image we did what we like to call quasi-LIC: injecting a
high amount of depth-sorted, semi-transparent particles yields stream-like structures which intensity corresponds to particle density. The difference to traditional LIC is
that the structures are automatically thinned out to interesting regions.
[TOP]
Visualization Performed 2
 In this image we show the DTI brain data set rendered using stream tubes. The standard color-mapping scheme based on fractional anisotropy (FA) was used.
From the probe fibers emerge as they are traced along the tensors' diffusion directions. Important brain structures can be easily destinguished in this image:
the superior longitudinal fasciculus is colored in green, parts of the corpus callosum can be made out in red, and the dark blue region to the left, outside the
probe is the corona radiata. Sadly the data set is rather noisy, but a demonstration of the particle engine with high quality data sets can be found in our
IEEE Visualization 2005 paper.
In this image we show the DTI brain data set rendered using stream tubes. The standard color-mapping scheme based on fractional anisotropy (FA) was used.
From the probe fibers emerge as they are traced along the tensors' diffusion directions. Important brain structures can be easily destinguished in this image:
the superior longitudinal fasciculus is colored in green, parts of the corpus callosum can be made out in red, and the dark blue region to the left, outside the
probe is the corona radiata. Sadly the data set is rather noisy, but a demonstration of the particle engine with high quality data sets can be found in our
IEEE Visualization 2005 paper.
This visualization renders at ca. 5fps on our system described above.
[TOP]
Additional Comments
The presented particle engine is well suited to the visualization of 3D flow, tensors, and multi-modal data in general. Despite its great flexibility
and performance there remain some issues, though. For performance reasons we can only handle regular cartesian or rectilinear grids. We are, however, investigating
ways to integrate multiblock grids into the engine. Another issue is the limited amount of GPU memory. We are investigating compression schemes to alleviate
this barrier, but on the other hand there are already GPUs with 512MB RAM available, and 1GB GPU memory is just a matter of (short) time. The reward for
the arrangement with these issues is the fact that the rendering performance scales with GPU power, not with the CPU. Tests on a nVidia 7800GTX indicate
a performance gain of roughly a factor of two, which fits nicely into the theory that GPUs more than double their performance every year.
Note that all images can be clicked to obtain higher resolution images. Some of the images in Task 3 were downsampled from 1600x1200 due to size restrictions.
Feel free to contact us to obtain the original images.
We are sorry for the need to resort to the wmv files, but since particles tend to get really eaten up by the codec, we chose wmv since it produced better results than DivX.
|
 |